𝗧𝗵𝗲 𝗠𝗮𝘁𝗵𝗲𝗺𝗮𝘁𝗶𝗰𝗮𝗹 𝗔𝗿𝗰𝗵𝗶𝘁𝗲𝗰𝘁𝘂𝗿𝗲 𝗼𝗳 𝗡𝗲𝘂𝗿𝗼𝗽𝗹𝗮𝘀𝘁𝗶𝗰𝗶𝘁𝘆: 𝗜𝗻𝘁𝗲𝗴𝗿𝗮𝘁𝗶𝗻𝗴 𝗗𝗘 𝗠𝗼𝗱𝗲𝗹𝘀 𝘄𝗶𝘁𝗵 𝗠𝗼𝗱𝗲𝗿𝗻 𝗠𝗟. 𝗕𝗲𝘆𝗼𝗻𝗱 𝘁𝗵𝗲 𝗦𝗶𝗻𝗴𝗹𝗲 𝗡𝗲𝘂𝗿𝗼𝗻: 𝗔 𝗨𝗻𝗶𝗳𝗶𝗲𝗱 𝗛𝗶𝗲𝗿𝗮𝗿𝗰𝗵𝘆 𝗳𝗼𝗿 𝗠𝘂𝗹𝘁𝗶𝘀𝗰𝗮𝗹𝗲 𝗡𝗲𝘂𝗿𝗮𝗹 𝗠𝗼𝗱𝗲𝗹𝗶𝗻𝗴. 𝗕𝗿𝗶𝗱𝗴𝗶𝗻𝗴 𝘁𝗵𝗲 𝗚𝗮𝗽: 𝗙𝗿𝗼𝗺 𝗗𝗲𝘁𝗲𝗿𝗺𝗶𝗻𝗶𝘀𝘁𝗶𝗰 𝗢𝗗𝗘𝘀 𝘁𝗼 𝗦𝘁𝗼𝗰𝗵𝗮𝘀𝘁𝗶𝗰 𝗣𝗼𝗽𝘂𝗹𝗮𝘁𝗶𝗼𝗻 𝗗𝘆𝗻𝗮𝗺𝗶𝗰𝘀.
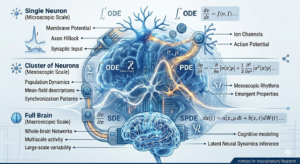
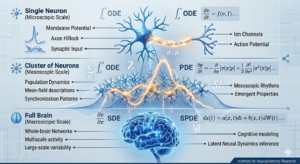
Large neuronal networks exhibit complex dynamics across multiple scales, from single-neuron excitability to whole-brain rhythms. At the Institute, we are examining how a unified hierarchy of differential equations can bridge these gaps. This framework connects deterministic, stochastic, and mean-field descriptions, providing a robust toolkit for multiscale modeling in computational neuroscience.
Ordinary Differential Equation (ODE) models, such as conductance-based systems, allow us to summarize macroscopic neural behavior through reduced variables. However, to understand population-level activity, we must transition to mean-field Partial Differential Equation (PDE) models. Equations like the Fokker-Planck or age-structured kinetic equations describe how probability densities evolve over synaptic states, linking individual mechanisms to collective oscillations.
Because variability is a hallmark of biological neural systems, our current focus emphasizes Stochastic Differential Equations (SDEs) and their extensions into jump-diffusion processes. These stochastic models are essential for describing random membrane fluctuations and irregular spike trains. They are not merely theoretical; they are critical for quantifying noise in electrophysiological recordings and inferring latent neural dynamics.
The versatility of this ODE-PDE-SDE framework offers a path toward integrated neural signal processing and cognitive modeling. By relating stochastic variability back to surrounding deterministic frameworks, we can better analyze bifurcations and collective patterns that define healthy versus pathological brain states.
We conclude by looking toward the next frontier: the integration of these differential equation models with modern machine learning. Addressing open challenges in multiscale inference and control-oriented formulations is essential for the future of neuroplasticity research and the development of advanced neuro-therapeutics.



 Posted in
Posted in